Report Displayers Examples¶
Report displayers you can use in your own Mine and some examples created for specific data types in modMine, FlyMine and metabolicMine.
The following displayers can all be used for data loaded by standard InterMine parsers. To see how to configure them check out FlyMine’s webconfig-model.xml.
SequenceFeature summary¶
Applicable for any SequenceFeature - shows length, sequence export, chromosome location, cyto location and SO term (where present).
<reportdisplayer javaClass="org.intermine.bio.web.displayer.SequenceFeatureDisplayer"
jspName="model/sequenceFeatureDisplayer.jsp"
replacesFields="chromosome,chromosomeLocation,sequence,length,sequenceOntologyTerm,locations,cytoLocation"
placement="summary"
types="SequenceFeature"/>

A Sequence feature displayer in metabolicMine.
Protein sequence¶
Applicable for Protein - shows length, sequence export.
<reportdisplayer javaClass="org.intermine.bio.web.displayer.ProteinSequenceDisplayer"
jspName="model/proteinSequenceDisplayer.jsp"
replacesFields="sequence"
placement="summary"
types="Protein"/>

A Protein sequence displayer in FlyMine.
GBrowse¶
Show an inline image from a configured GBrowse instance.
<reportdisplayer javaClass="org.intermine.bio.web.displayer.GBrowseDisplayer"
jspName="model/gbrowseDisplayer.jsp"
replacesFields=""
placement="Genes"
types="SequenceFeature"/>
This also needs two properties to be configured in the minename.properties file: gbrowse.prefix and gbrowse_image.prefix which give the location of a running GBrowse instance.
gbrowse.prefix=http://www.flymine.org/cgi-bin/gbrowse
gbrowse_image.prefix=http://www.flymine.org/cgi-bin/gbrowse_img
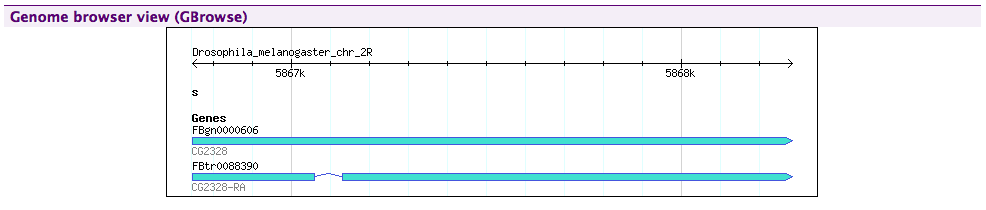
A Genome browser view in FlyMine.
Homologues¶
Shows a table of organism and homologous genes of homologues per organism.
<reportdisplayer javaClass="org.intermine.bio.web.displayer.HomologueDisplayer"
jspName="model/homologueDisplayer.jsp"
replacesFields="homologues"
placement="Homology"
parameters="{'dataSets': ['TreeFam data set', 'KEGG orthologues data set']}"
types="Gene"/>
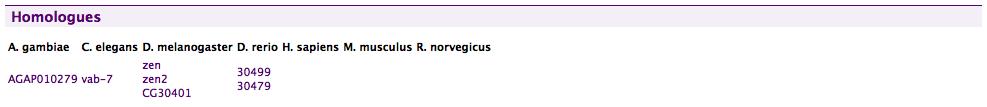
A Homologues displayer in FlyMine.
Note that FlyMine includes a specific displayer to show the twelve Drosophila species as a phylogenetic tree.
Gene structure¶
Displays transcripts, exons, introns, UTRs and CDSs if present in the model and for the particular organism. Can be added to report pages for any of these feature types and will find the parent gene and show all transcripts, highlighting the feature of the actual report page.
<reportdisplayer javaClass="org.intermine.bio.web.displayer.GeneStructureDisplayer"
jspName="model/geneStructureDisplayer.jsp"
replacesFields="transcripts,exons,CDSs,introns,UTRs,fivePrimeUTR,threePrimeUTR"
placement="Genomics"
types="Gene,Transcript,Exon,Intron,UTR,CDS"/>
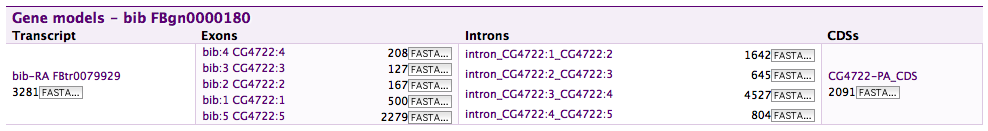
A Gene structure displayer in FlyMine.
Gene Ontology¶
Simple display of GO terms and evidence codes for a gene, grouped by branch in the ontology. Groups by the three main ontologies (function, process and component) so you may need to run the GO source.
<reportdisplayer javaClass="org.intermine.bio.web.displayer.GeneOntologyDisplayer"
jspName="model/geneOntologyDisplayer.jsp"
replacesFields="goAnnotation,ontologyAnnotations"
placement="Function"
types="Gene"/>

A Gene ontology displayer in modMine.
UniProt comments¶
A clear view of curated curated comments from UniProt (SwissProt) applied to a protein, or for a gene will show comments from all proteins of the gene.
<reportdisplayer javaClass="org.intermine.bio.web.displayer.UniProtCommentsDisplayer"
jspName="model/uniProtCommentsDisplayer.jsp"
replacesFields=""
placement="summary"
types="Gene,Protein"/>

A Uniprot curated comments displayer in metabolicMine.
Interaction network¶
Uses the Cytoscape Web plugin to display physical and genetics interactions. The interaction displayer links to report pages, allows creation of a gene list of the whole network and can show tabular interaction data. Read NetworkDisplayer for details.
<reportdisplayer javaClass="org.intermine.bio.web.displayer.CytoscapeNetworkDisplayer"
jspName="model/cytoscapeNetworkDisplayer.jsp"
replacesFields="interactions"
placement="Interactions"
types="Gene,Protein"/>
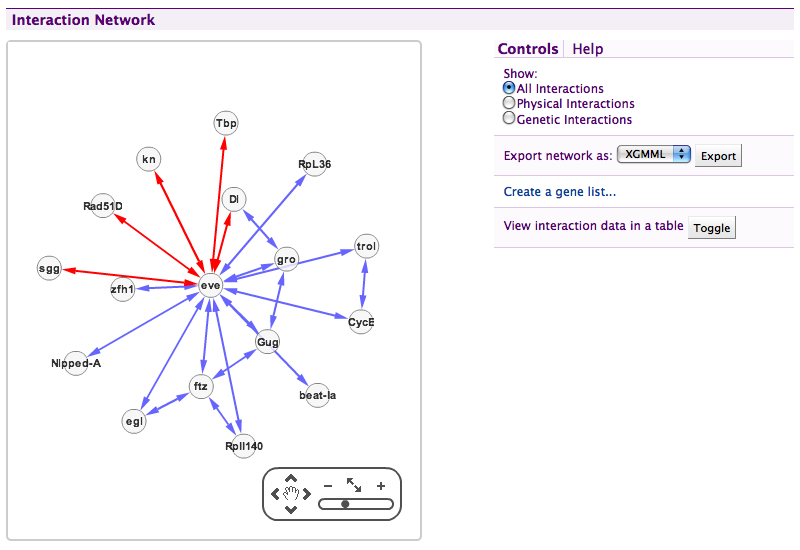
An Interactions displayer in FlyMine.
Overlapping features¶
A summary view of features that overlap the chromosome location of the reported feature, if the gene structure displayer is also used it will exclude any features that are part of the same gene model - i.e. it won’t report that a gene overlaps it’s own exons.
<reportdisplayer javaClass="org.intermine.bio.web.displayer.OverlappingFeaturesDisplayer"
jspName="model/overlappingFeaturesDisplayer.jsp"
replacesFields="overlappingFeatures"
placement="Genomics"
types="SequenceFeature"/>
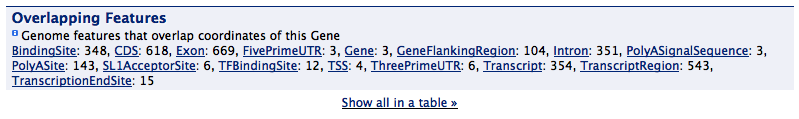
An Overlapping features displayer in modMine.
Complexes - Protein interactions¶
Viewer displaying complex interactions. Data must be loaded from IntAct. Original Source: http://interactionviewer.org/.
<reportdisplayer javaClass="org.intermine.bio.web.displayer.ComplexDisplayer"
jspName="model/complexDisplayer.jsp"
replacesFields=""
placement="summary"
types="Complex"/>
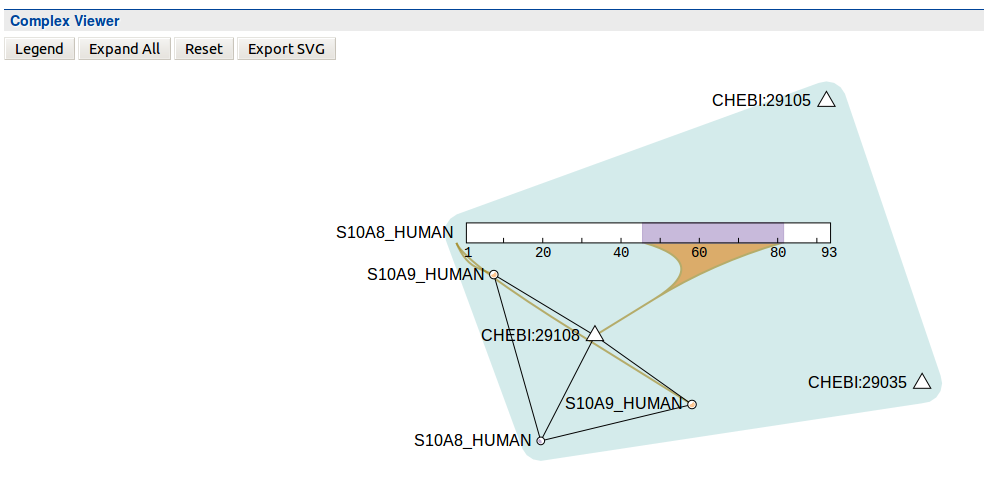

A Complex interaction displayer in HumanMine.
Specific Displayers¶
There are some displayers created for specific data sets in FlyMine, metabolicMine or modMine that may not be re-usable in other Mines but could be adapted or provide inspiration.
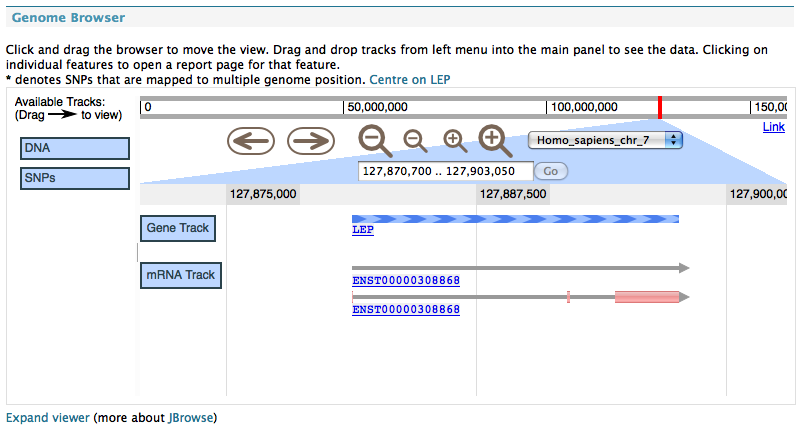
JBrowse genome browser in metabolicMine.
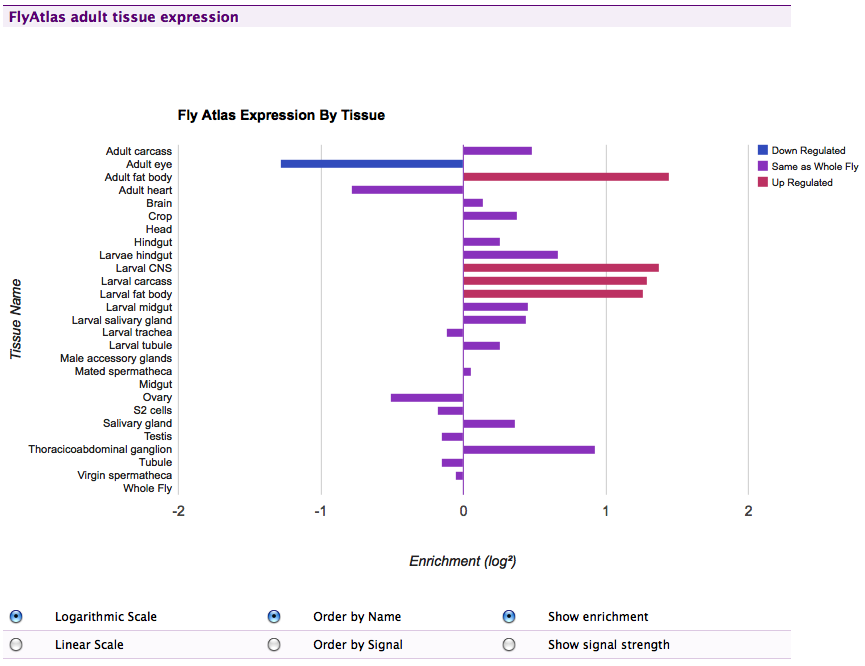
FlyAtlas gene experssion data in FlyMine, this uses the Google Data Vizualization API JavaScript library to render an interactive graph in the browser.
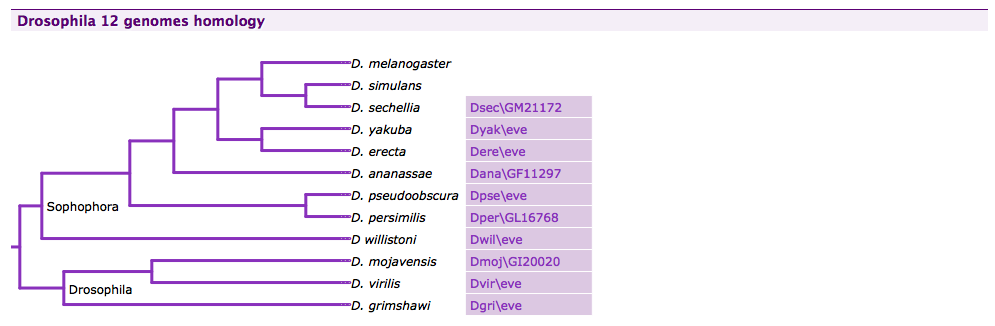
A phylogenetic tree of Drosophila species displayed using the jsPhyloSVG JavaScript library in FlyMine.
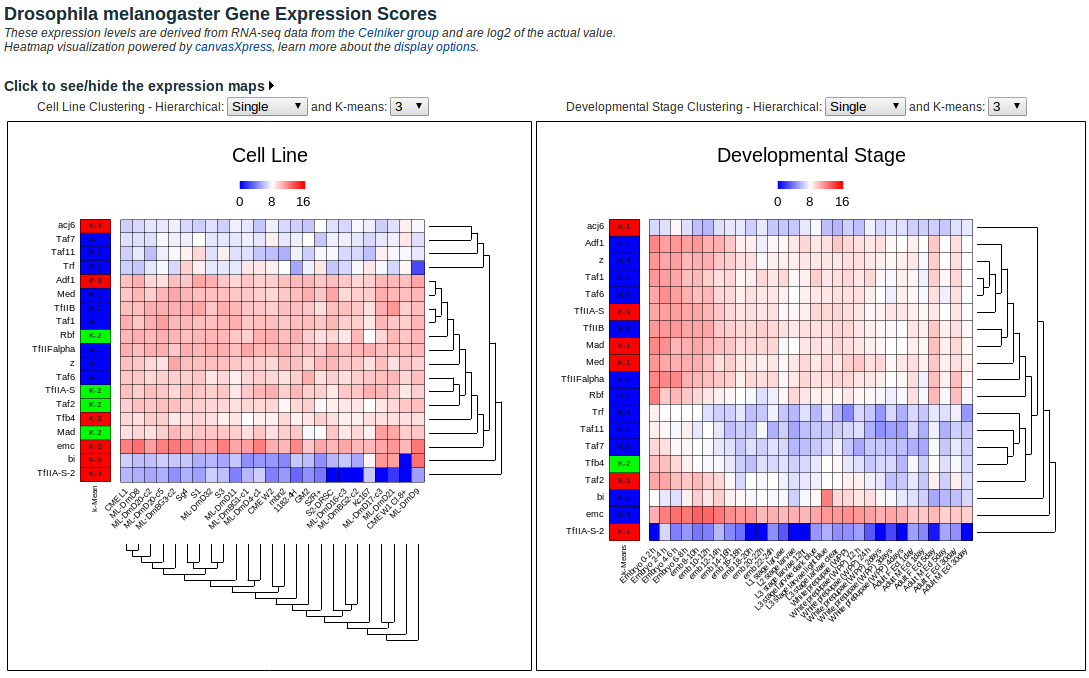
Heatmap of fly gene expression in modMine, this makes use of canvasXpress JavaScript library.
